Structural search and advanced query search is temporarily unavailable. We are working to fix this issue. Thank you for your support and patience.
L-Idonate (M2MDB001772)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-07-30 14:55:31 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-06-04 16:52:57 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | L-Idonate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | L-idonate is a member of the chemical class known as Sugar Acids and Derivatives. These are compounds containing a saccharide unit which bears a carboxylic acid group. L-idonate is a metabolite in D-gluconate catabolism. The catabolic sequence for l-idonate is as follows: l-idonate is transported by the l-idonate transporter, IdnT; l-idonate is oxidized to 5-keto-gluconate (5KG) by l-idonate 5-dehydrogenase, IdnD; 5KG is reduced to d-gluconate by 5-keto-D-gluconate 5-reductase, IdnO; and d-gluconate is phosphorylated by a thermosensitive gluconate kinase, IdnK, to make 6-phosphogluconate (6PG), which is further catabolized via the Entner-Doudoroff pathway. Thus, IdnD and IdnO allow for the redox-coupled interconversion of l-idonate to d-gluconate via 5KG. The l-idonate catabolic pathway overlaps d-gluconate catabolism through the common intermediates d-gluconate and 6PG. (PMID: 14973046) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
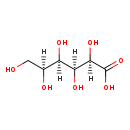
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C6H12O7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 196.1553 Monoisotopic: 196.058302738 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | RGHNJXZEOKUKBD-SKNVOMKLSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C6H12O7/c7-1-2(8)3(9)4(10)5(11)6(12)13/h2-5,7-11H,1H2,(H,12,13)/t2-,3+,4-,5+/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 1114-17-6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2R,3S,4R,5S)-2,3,4,5,6-pentahydroxyhexanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | L-idonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | [H][C@](O)(CO)[C@@]([H])(O)[C@]([H])(O)[C@@]([H])(O)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as medium-chain hydroxy acids and derivatives. These are hydroxy acids with a 6 to 12 carbon atoms long side chain. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Hydroxy acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Medium-chain hydroxy acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Medium-chain hydroxy acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | L-Idonate + NADP <> 5-Dehydro-D-gluconate + NADPH + Hydrogen ion NAD(P)<sup>+</sup> + L-Idonate <> NAD(P)H + 5-Keto-D-gluconate + Hydrogen ion 2-keto-L-gulonate + NAD(P)H + Hydrogen ion > L-Idonate + NAD(P)<sup>+</sup> Hydrogen ion + 2-Dehydro-D-gluconate + NADPH <> L-Idonate + NADP L-Idonate + NAD(P)(+) > 5-Keto-D-gluconate + NAD(P)H L-Idonate + NAD + NADP <> 5-Keto-D-gluconate + NADH + NADPH + Hydrogen ion 2-Keto-L-gluconate + Hydrogen ion + NADPH + 2-Dehydro-D-gluconate + NADPH > NADP + L-Idonate L-Idonate + NADP > Hydrogen ion + NADPH + 5-Keto-D-gluconate + NADPH + 5-Keto-D-gluconate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor
- Specific function:
- Catalyzes the NADPH-dependent reduction of glyoxylate and hydroxypyruvate into glycolate and glycerate, respectively. Can also reduce 2,5-diketo-D-gluconate (25DKG) to 5-keto-D- gluconate (5KDG), 2-keto-D-gluconate (2KDG) to D-gluconate, and 2- keto-L-gulonate (2KLG) to L-idonate (IA), but it is not its physiological function. Inactive towards 2-oxoglutarate, oxaloacetate, pyruvate, 5-keto-D-gluconate, D-fructose and L- sorbose. Activity with NAD is very low
- Gene Name:
- ghrB
- Uniprot ID:
- P37666
- Molecular weight:
- 35395
Reactions
| Glycolate + NADP(+) = glyoxylate + NADPH. |
| D-glycerate + NAD(P)(+) = hydroxypyruvate + NAD(P)H. |
| D-gluconate + NADP(+) = 2-dehydro-D-gluconate + NADPH. |
- General function:
- Involved in zinc ion binding
- Specific function:
- Catalyzes the NADH/NADPH-dependent oxidation of L- idonate to 5-ketogluconate (5KG)
- Gene Name:
- idnD
- Uniprot ID:
- P39346
- Molecular weight:
- 37146
Reactions
| L-idonate + NAD(P)(+) = 5-dehydrogluconate + NAD(P)H. |
Transporters
- General function:
- Involved in gluconate transmembrane transporter activity
- Specific function:
- Transports L-idonate, D-gluconate and 5-keto-D- gluconate, from the periplasm across the inner membrane
- Gene Name:
- idnT
- Uniprot ID:
- P39344
- Molecular weight:
- 46041