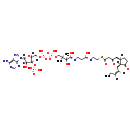| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2012-05-31 14:26:07 -0600 |
|---|
| Update Date | 2015-06-03 17:19:15 -0600 |
|---|
| Secondary Accession Numbers | |
|---|
| Identification |
|---|
| Name: | 3-Oxo-OPC4-CoA |
|---|
| Description | 3-oxo-opc4-coa belongs to the class of Coenzyme A and Derivatives. These are derivative of vitamin B5 containing a 4'-phosphopantetheine moiety attached to a diphospho-adenosine. (inferred from compound structure) |
|---|
| Structure | |
|---|
| Synonyms: | Not Available |
|---|
| Chemical Formula: | C35H54N7O19P3S |
|---|
| Weight: | Average: 1001.825
Monoisotopic: 1001.240802807 |
|---|
| InChI Key: | QGJLCXXJEFRWHP-ZRHHFSRKSA-N |
|---|
| InChI: | InChI=1S/C35H54N7O19P3S/c1-4-5-6-7-22-20(8-9-23(22)44)14-21(43)15-26(46)65-13-12-37-25(45)10-11-38-33(49)30(48)35(2,3)17-58-64(55,56)61-63(53,54)57-16-24-29(60-62(50,51)52)28(47)34(59-24)42-19-41-27-31(36)39-18-40-32(27)42/h5-6,18-20,22,24,28-30,34,47-48H,4,7-17H2,1-3H3,(H,37,45)(H,38,49)(H,53,54)(H,55,56)(H2,36,39,40)(H2,50,51,52)/b6-5-/t20-,22+,24-,28-,29-,30?,34-/m1/s1 |
|---|
| CAS number: | Not Available |
|---|
| IUPAC Name: | 4-({[({[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)oxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy](hydroxy)phosphoryl}oxy)-2-hydroxy-3,3-dimethyl-N-(2-{[2-({3-oxo-4-[(1R,2S)-3-oxo-2-[(2Z)-pent-2-en-1-yl]cyclopentyl]butanoyl}sulfanyl)ethyl]-C-hydroxycarbonimidoyl}ethyl)butanimidic acid |
|---|
| Traditional IUPAC Name: | 3-Oxo-OPC4-CoA |
|---|
| SMILES: | CC\C([H])=C(\[H])C[C@]1([H])C(=O)CC[C@]1([H])CC(=O)CC(=O)SCCN=C(O)CCN=C(O)C(O)([H])C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@@]1([H])O[C@@]([H])(N2C=NC3=C(N)N=CN=C23)[C@@](O)([H])[C@]1([H])OP(O)(O)=O |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 3-oxo-acyl coas. These are organic compounds containing a 3-oxo acylated coenzyme A derivative. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
| Direct Parent | 3-oxo-acyl CoAs |
|---|
| Alternative Parents | |
|---|
| Substituents | - Coenzyme a or derivatives
- Purine ribonucleoside 3',5'-bisphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside diphosphate
- Ribonucleoside 3'-phosphate
- Pentose phosphate
- Pentose-5-phosphate
- Beta amino acid or derivatives
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Monosaccharide phosphate
- Organic pyrophosphate
- Pentose monosaccharide
- Imidazopyrimidine
- Purine
- Monoalkyl phosphate
- Aminopyrimidine
- Alkyl phosphate
- 1,3-dicarbonyl compound
- Imidolactam
- Monosaccharide
- N-acyl-amine
- N-substituted imidazole
- Organic phosphoric acid derivative
- Pyrimidine
- Fatty amide
- Phosphoric acid ester
- Tetrahydrofuran
- Imidazole
- Heteroaromatic compound
- Azole
- Cyclic ketone
- Carbothioic s-ester
- Secondary alcohol
- Ketone
- Carboxamide group
- Thiocarboxylic acid ester
- Amino acid or derivatives
- Secondary carboxylic acid amide
- Sulfenyl compound
- Thiocarboxylic acid or derivatives
- Organoheterocyclic compound
- Azacycle
- Oxacycle
- Carboxylic acid derivative
- Organosulfur compound
- Organic oxygen compound
- Hydrocarbon derivative
- Carbonyl group
- Organic nitrogen compound
- Primary amine
- Organopnictogen compound
- Organic oxide
- Organooxygen compound
- Organonitrogen compound
- Amine
- Alcohol
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Physical Properties |
|---|
| State: | Not Available |
|---|
| Charge: | -3 |
|---|
| Melting point: | Not Available |
|---|
| Experimental Properties: | |
|---|
| Predicted Properties | |
|---|
| Biological Properties |
|---|
| Cellular Locations: | Cytoplasm |
|---|
| Reactions: | |
|---|
| SMPDB Pathways: | Not Available |
|---|
| KEGG Pathways: | - alpha-Linolenic acid metabolism ec00592
|
|---|
| EcoCyc Pathways: | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| Spectra |
|---|
| Spectra: | |
|---|
| References |
|---|
| References: | - Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. (2012). "KEGG for integration and interpretation of large-scale molecular data sets." Nucleic Acids Res 40:D109-D114. Pubmed: 22080510
|
|---|
| Synthesis Reference: | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| Links |
|---|
| External Links: | | Resource | Link |
|---|
| CHEBI ID | 80459 | | HMDB ID | Not Available | | Pubchem Compound ID | 23724725 | | Kegg ID | C16338 | | ChemSpider ID | Not Available | | Wikipedia ID | Not Available | | BioCyc ID | Not Available |
|
|---|