| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2012-05-31 13:53:57 -0600 |
|---|
| Update Date | 2015-06-03 15:54:10 -0600 |
|---|
| Secondary Accession Numbers | |
|---|
| Identification |
|---|
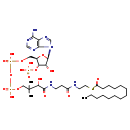
| Name: | Tetradecanoyl-CoA |
|---|
| Description | Tetradecanoyl-CoA (or myristoyl-CoA) is an intermediate in fatty acid biosynthesis, fatty acid elongation and the beta oxidation of fatty acids. It is also used in the myristoylation of proteins. The first pass through the beta-oxidation process starts with the saturated fatty acid palmitoyl-CoA and produces myristoyl-CoA. A total of four enzymatic steps are required, starting with VLCAD CoA dehydrogenase (Very Long Chain) activity, followed by three enzymatic steps catalyzed by enoyl-CoA hydratase, 3-hydroxyacyl-CoA dehydrogenase, and ketoacyl-CoA thiolase. Myristoylation of proteins is also catalyzed by the presence of myristoyl-CoA along with Myristoyl-CoA:protein N-myristoyltransferase (NMT). Myristoylation is an irreversible, co-translational (during translation) protein modification found in animals, plants, fungi and viruses. In this protein modification a myristoyl group (derived from myristioyl CoA) is covalently attached via an amide bond to the alpha-amino group of an N-terminal amino acid of a nascent polypeptide. It is more common on glycine residues but also occurs on other amino acids. Myristoylation also occurs post-translationally, for example when previously internal glycine residues become exposed by caspase cleavage during apoptosis. |
|---|
| Structure | |
|---|
| Synonyms: | - Myristoyl-CoA
- Myristoyl-coenzyme A
- N-C14:0CoA
- N-C14:0Coenzyme A
- S-Tetradecanoyl-coenzyme A
- Tetradecanoyl CoA
- Tetradecanoyl Coenzyme A
- Tetradecanoyl-CoA
- Tetradecanoyl-CoA (N-C14:0CoA)
- Tetradecanoyl-coenzyme A
|
|---|
| Chemical Formula: | C35H62N7O17P3S |
|---|
| Weight: | Average: 977.89
Monoisotopic: 977.313573819 |
|---|
| InChI Key: | DUAFKXOFBZQTQE-XVDJLSDJSA-N |
|---|
| InChI: | InChI=1S/C35H62N7O17P3S/c1-4-5-6-7-8-9-10-11-12-13-14-15-26(44)63-19-18-37-25(43)16-17-38-33(47)30(46)35(2,3)21-56-62(53,54)59-61(51,52)55-20-24-29(58-60(48,49)50)28(45)34(57-24)42-23-41-27-31(36)39-22-40-32(27)42/h22-24,28-30,34,45-46H,4-21H2,1-3H3,(H,37,43)(H,38,47)(H,51,52)(H,53,54)(H2,36,39,40)(H2,48,49,50)/t24-,28-,29-,30?,34-/m1/s1 |
|---|
| CAS number: | 3130-72-1 |
|---|
| IUPAC Name: | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-2-({[hydroxy({[hydroxy({3-hydroxy-2,2-dimethyl-3-[(2-{[2-(tetradecanoylsulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]propoxy})phosphoryl]oxy})phosphoryl]oxy}methyl)oxolan-3-yl]oxy}phosphonic acid |
|---|
| Traditional IUPAC Name: | [(2R,3S,4R,5R)-5-(6-aminopurin-9-yl)-4-hydroxy-2-[({hydroxy[hydroxy(3-hydroxy-2,2-dimethyl-3-[(2-{[2-(tetradecanoylsulfanyl)ethyl]carbamoyl}ethyl)carbamoyl]propoxy)phosphoryl]oxyphosphoryl}oxy)methyl]oxolan-3-yl]oxyphosphonic acid |
|---|
| SMILES: | CCCCCCCCCCCCCC(=O)SCCNC(=O)CCNC(=O)C(O)C(C)(C)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1OP(O)(O)=O)N1C=NC2=C1N=CN=C2N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as long-chain fatty acyl coas. These are acyl CoAs where the group acylated to the coenzyme A moiety is a long aliphatic chain of 13 to 21 carbon atoms. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl thioesters |
|---|
| Direct Parent | Long-chain fatty acyl CoAs |
|---|
| Alternative Parents | |
|---|
| Substituents | - Coenzyme a or derivatives
- Purine ribonucleoside 3',5'-bisphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside diphosphate
- Ribonucleoside 3'-phosphate
- Pentose phosphate
- Pentose-5-phosphate
- Beta amino acid or derivatives
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Monosaccharide phosphate
- Organic pyrophosphate
- Pentose monosaccharide
- Imidazopyrimidine
- Purine
- Monoalkyl phosphate
- Aminopyrimidine
- Imidolactam
- N-acyl-amine
- N-substituted imidazole
- Organic phosphoric acid derivative
- Monosaccharide
- Pyrimidine
- Alkyl phosphate
- Fatty amide
- Phosphoric acid ester
- Tetrahydrofuran
- Imidazole
- Azole
- Heteroaromatic compound
- Carbothioic s-ester
- Secondary alcohol
- Thiocarboxylic acid ester
- Carboxamide group
- Secondary carboxylic acid amide
- Amino acid or derivatives
- Sulfenyl compound
- Thiocarboxylic acid or derivatives
- Organoheterocyclic compound
- Azacycle
- Oxacycle
- Carboxylic acid derivative
- Organosulfur compound
- Organic oxygen compound
- Hydrocarbon derivative
- Carbonyl group
- Organic nitrogen compound
- Primary amine
- Organopnictogen compound
- Organic oxide
- Organooxygen compound
- Organonitrogen compound
- Alcohol
- Amine
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Physical Properties |
|---|
| State: | Solid |
|---|
| Charge: | -4 |
|---|
| Melting point: | Not Available |
|---|
| Experimental Properties: | |
|---|
| Predicted Properties | |
|---|
| Biological Properties |
|---|
| Cellular Locations: | Cytoplasm |
|---|
| Reactions: | |
|---|
| SMPDB Pathways: | |
|---|
| KEGG Pathways: | |
|---|
| EcoCyc Pathways: | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| Spectra |
|---|
| Spectra: | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-1911101102-349a13615858f61c2639 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0941300000-9797f6c398276f5d5e98 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-2900102000-cc4f9c8978927acec695 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a7i-4971342506-ab177bf279e0e32e2b12 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0560-3921201001-624d08d1bbb66df93c2e | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-057i-6900000000-d973a8b2d469b04161eb | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0000000009-3239f04ec07dfb05c844 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-4120302409-5222d96c1a16a7c2a7df | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0570-2102301209-fd8e6272e04594f37eaa | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004i-0000000009-7a4d637e9994bab811eb | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0800100449-955bc85312168f96efe1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00di-0101900000-c9607999f9d61657c1fb | View in MoNA |
|---|
| MS | Mass Spectrum (Electron Ionization) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum | Not Available | View in JSpectraViewer |
|---|
|
|---|
| References |
|---|
| References: | - Hunt MC, Ruiter J, Mooyer P, van Roermond CW, Ofman R, Ijlst L, Wanders RJ: Identification of fatty acid oxidation disorder patients with lowered acyl-CoA thioesterase activity in human skin fibroblasts. Eur J Clin Invest. 2005 Jan;35(1):38-46. Pubmed: 15638818
- Kanehisa, M., Goto, S., Sato, Y., Furumichi, M., Tanabe, M. (2012). "KEGG for integration and interpretation of large-scale molecular data sets." Nucleic Acids Res 40:D109-D114. Pubmed: 22080510
- Keseler, I. M., Collado-Vides, J., Santos-Zavaleta, A., Peralta-Gil, M., Gama-Castro, S., Muniz-Rascado, L., Bonavides-Martinez, C., Paley, S., Krummenacker, M., Altman, T., Kaipa, P., Spaulding, A., Pacheco, J., Latendresse, M., Fulcher, C., Sarker, M., Shearer, A. G., Mackie, A., Paulsen, I., Gunsalus, R. P., Karp, P. D. (2011). "EcoCyc: a comprehensive database of Escherichia coli biology." Nucleic Acids Res 39:D583-D590. Pubmed: 21097882
|
|---|
| Synthesis Reference: | Not Available |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| Links |
|---|
| External Links: | |
|---|